
Concept explainers
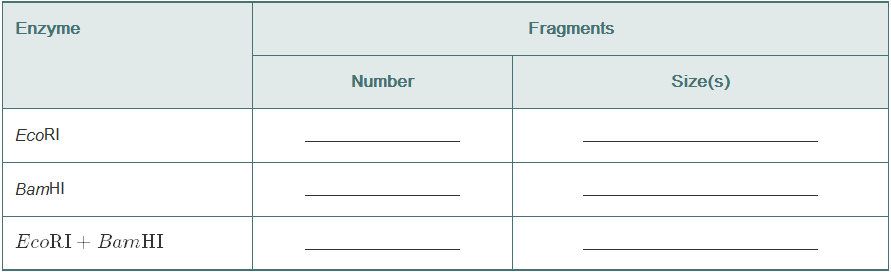
Using this map of pLAB30 (18 kb), give the number and lengths of the restriction fragments that would result from digesting pLAB30 with EcoR1, with BamHI, and with both enzymes together.

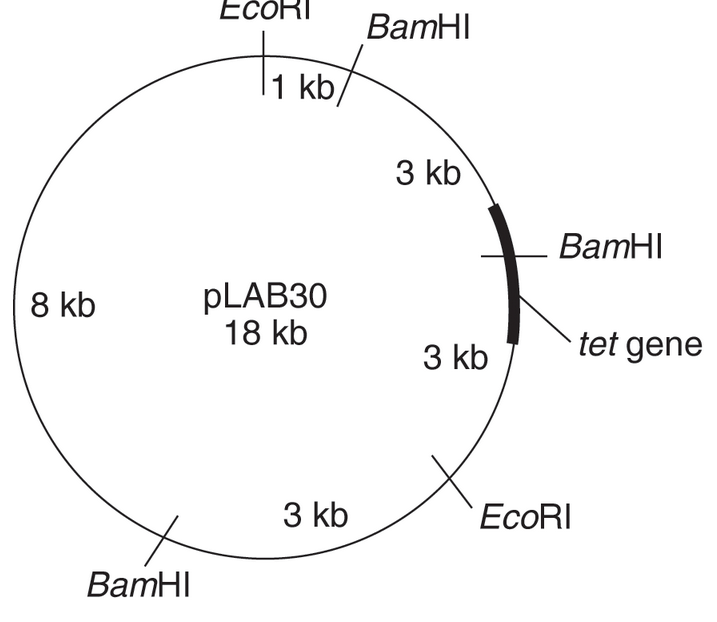
Which enzyme will give the smallest piece containing the tetracycline-resistance gene?
To determine:
The size and number of DNA Fragments given by restriction enzymes Eco RI, Bam HI, and both these enzymes together.
Introduction:
Restriction enzyme is an enzyme that cuts the DNA fragment at the specific site by recognizing the DNA sequence at that site. The site at which the DNA is cut is termed as the restriction site.
Explanation of Solution
pLAB30 is an 18 kb map, which is cut by two restriction enzymes, Eco RI, and Bam HI. When Eco RI acts, then it cuts the DNA fragment at two sites, giving an 11 kb fragment and 7 kb fragment of the DNA.
When Bam HIacts, then it cuts DNA fragment at three sites, giving three fragments. The size of these fragments is 9 kb, 3 kb, and 6 kb.
When both these restriction enzymes act together, then 5 DNA fragments are obtained. The size of these fragments is 1 kb, 3 kb, 3 kb, 3kb, and 8 kb.
The table showing the size and number of DNA fragments after the DNA is acted upon by restriction enzymes is shown below:
| Restriction enzyme | Number of DNA fragments obtained | Size of DNA fragments |
| Eco RI | 2 | 11 kb and 7 kb |
| Bam HI | 3 | 9 kb, 3 kb, and 6 kb |
| Eco RI + Bam HI | 5 | 1 kb, 3 kb, 3 kb, 3kb, and 8 kb |
Want to see more full solutions like this?
Chapter 30 Solutions
Laboratory Experiments in Microbiology (12th Edition) (What's New in Microbiology)
- Mutation analysis of GCK gene in patients with diabetes revealed a c.114 T->A (shown in bold and underlined) substitution in heterozygote state. In order to check the mutation in healthy individuals, restriction enzyme analysis will be used. a) which enzyme can we use to differentiate wild type and mutant sequence? Please indicate which allele (wild type or mutant allele) will be cut with the restriction enzyme. Use table 1 shown below. b) ATGAGGCTCTTTGCCACCAGTCCCAGTTTTATGCATGGCAGCTCTAATGACAGGATGGTCACCCCTG СTGAGGCCACTCCTGGTCACCATGACAАССАCAGGCCCTCTТСAGTATCACAGTAAGCCCTGGCAGG AGAATCCCCCACTCCACACCTGGCTGGAGCACGAAATGCCGAGCGGCGCCTGAGCCCCAGGGAAG CAGGCTAGGATGTGA Figure 1. GCK gene sequence. Length of the fragment is 213bp. Table1. The restriction enzymes and their recognition sequences. Bestriction enzyme Recognition sequence Nari GG/CGCC Ddel C/TOAG Hae II DGCGC/n Hpal cc/GG Alul AG/CT Smal ccc/GGG Mbol /GATC Mae II IGTDAC Bsp 1286 I GNGCn/c Hind II A/AGCTT ECOR I G/AATTC D: any Ducleotide 1:…arrow_forwardMutation analysis of GCK gene in patients with diabetes revealed a c.114 T→A (shown in bold and underlined) substitution in heterozygote state. In order to check the mutation in healthy individuals, restriction enzyme analysis will be used. a) Which enzyme can we use to differentiate wild type and mutant sequence? Please indicate which allele (wild type or mutant allele) will be cut with the restriction enzyme. Use table 1 shown below. b) Draw the expected agarose gel result of a homozygous wild type, homozygous mutant and heterozygote individual after restriction enzyme analysis. ATGAGGCTCTTTGCCACCAGTCCCAGTTTTATGCATGGCAGCTCTAATGACAGGATGGTCACCCCTGCTGAGGCC ACTCCTGGTCACCATGACAACCACAGGCCCTCTCAGTATCACAGTAAGCCCTGGCAGGAGAATCCCCCACTCCAC ACCTGGCTGGAGCACGAAATGCCGAGCGGCGCCTGAGCCCCAGGGAAGCAGGCTAGGATGTGA Figure 1. GCK gene sequence. Length of the fragment is 213bp. Table1. The restriction enzymes and their recognition sequences. Restriction enzyme Recognition seguence www wwwtw ww Nar I GG/CGCC…arrow_forwardAssume that a circular plasmid is 3200 base pairs in length and has restriction sites for HindIII restriction enzyme at the following locations: 400, 700, 1400, 2600. Give the expected sizes of the restriction fragments following complete digestion.arrow_forward
- Knowing that you are using HindIII and EcoRI to cut your plasmids, and that those two enzymes cut within the MCS, use the map of pUC19 provided below to compute: What will be the sizes of the 2 restriction fragments if NO insert is present in pUC19? What will be the sizes of the 2 restriction fragments if the approximately 317 bp RT-PCR product (insert from WT satC dimer) was ligated successfully into the SmaI site? What will be the sizes of the 2 restriction fragments if TWO approximately 317 bp RT-PCR products (2 ligated inserts from WT satC dimer) were ligated into the SmaI site?arrow_forwardIf the sequence of base pairs along a DNA molecule occurs strictly at random, what is the expected frequency of a specific restriction enzyme recognition sequence of length (a) four and (b) six base pairs?arrow_forwardFor the DNA sequence shown, indicate the products of its cleavage with the following restriction endonucleases (AKA restriction enzymes):5′-ACAGCTGATTCGAATTCACGTT-3′3′-TGTCGACTAAGCTTAAGTGCAA-5′a) EcoRI (the recognition sequence and cleavage site is G↓AATTC);b) AluI (the recognition sequence and cleavage site is AG↓CT).arrow_forward
- Examine the structure of the pBR322 plasmid depicted below. Assume total size of the plasmid is 4,361 bp and the blue numbers indicate locations of restriction sites relative to the O point at the top of the plasmid. What size fragments would be generated by the following restriction digestion reactions? 1. Sall 2. Sal 1 + BamH1 3. Sal 1 + EcoR1 4. Sal I + BamH1 + EcoR1 PstI 3607 3000 4000 amp ori HindIII Edit View Insert Format Tools Table EcoRI EcoRV 4359 0 29 185 pBR322 4361 bp 2295 NdeI tet 2000 BamHI 375 651 SalI 1000 Type a short answer in the space provided below.arrow_forwardGiven the following double-stranded fragment of DNA: 5'- ACTTGGCAGGCCTTCGATCC-3' 3'- TGAАССGTCСGGAAGCTAGG-5' A hypothetical restriction endonuclease recognizes a 6bp sequence with two-fold symmetry (typical for restriction enzymes) found in this fragment and catalyzes cleavage of this DNA on both strands between GG nucleotides within the recognition sequence. This nuclease exhibits b-type cleavage (atypical for restriction enzymes). Draw the double-stranded sequence of each fragment after cleavage showing any phosphates left on the ends.arrow_forwardThe cellulase gene of Bacillus licheniformis was successfully cloned into the pET21a vector and expressed in Escherichia coli BL21. The pET21a vector consists of ampicillin resistant gene. To screen for the successful transformants, E. coli BL21 was cultivated on LB agar containing ampicillin (100 pg/mL) and 0.5% (w/v) carboxymethylcellulose, and incubated at 37 0C for 6 hours. After that the agar plate was stained with Congo red solution for 15 minutes and washed twice with sodium chloride solution and the observation is as shown in Figure 3. Answer the following: (i.) Briefly explain why ampicillin was added to the LB agar. (ii) Indicate the function of carboxymethylcellulose in the LB agar. (iii) Conclude how the researchers were able to identify the E. coli BL21 that carried the cellulase gene.arrow_forward
- Table 1 shows a list of restriction endonucleases with their recognition sequence and the sites of cleavage indicated by arrows. Table 1 Enzyme name Recognition sequence and position of cut 5'GIAATTC3 5'G!GATCC3' 5'GIGTACC3 5'GCIGGCCGC3' 5'IGATC3' 5'GGTACIC3' 5'ALGATCT3 EcoRI ВатHI Аcс651 Notl Sau3A Kpnl BglII (i) Which restriction enzyme(s) produce blunt ends? (ii) Are there any pair of neoschizomers in the list? Explain. (iii) Are there any pair of isocaudomers in the list? Explain.arrow_forwardA 12 kb linear DNA fragment is subject to single or double RE digest and agarose gelelectrophoresis, to yield the gel profile shown below. The first lane contains the size marker(M).a) Explain how the name of the enzyme EcoRI is derived.b) How many sites are there for EcoRI and PvuII respectively on this DNA fragment?c) Use the sizes of the DNA bands on the gel to compile a restriction enzyme map of the DNAfragment. Indicate the positions of the restriction enzymes sites for EcoRI and PvuII on themap.arrow_forwardThey have purified a 1100 bp HindIII restriction fragment that they plan to sequence. As a first step, they decided to construct a restriction map of the fragment for the enzymes EcoRI and SmaI. Below is shown an agarose gel of the appropriate digests. Draw a restriction map of the fragment and show the distances, in base pairs, between the HindIII, EcoRI, and SmaI sites. *arrow_forward
 Human Anatomy & Physiology (11th Edition)BiologyISBN:9780134580999Author:Elaine N. Marieb, Katja N. HoehnPublisher:PEARSON
Human Anatomy & Physiology (11th Edition)BiologyISBN:9780134580999Author:Elaine N. Marieb, Katja N. HoehnPublisher:PEARSON Biology 2eBiologyISBN:9781947172517Author:Matthew Douglas, Jung Choi, Mary Ann ClarkPublisher:OpenStax
Biology 2eBiologyISBN:9781947172517Author:Matthew Douglas, Jung Choi, Mary Ann ClarkPublisher:OpenStax Anatomy & PhysiologyBiologyISBN:9781259398629Author:McKinley, Michael P., O'loughlin, Valerie Dean, Bidle, Theresa StouterPublisher:Mcgraw Hill Education,
Anatomy & PhysiologyBiologyISBN:9781259398629Author:McKinley, Michael P., O'loughlin, Valerie Dean, Bidle, Theresa StouterPublisher:Mcgraw Hill Education, Molecular Biology of the Cell (Sixth Edition)BiologyISBN:9780815344322Author:Bruce Alberts, Alexander D. Johnson, Julian Lewis, David Morgan, Martin Raff, Keith Roberts, Peter WalterPublisher:W. W. Norton & Company
Molecular Biology of the Cell (Sixth Edition)BiologyISBN:9780815344322Author:Bruce Alberts, Alexander D. Johnson, Julian Lewis, David Morgan, Martin Raff, Keith Roberts, Peter WalterPublisher:W. W. Norton & Company Laboratory Manual For Human Anatomy & PhysiologyBiologyISBN:9781260159363Author:Martin, Terry R., Prentice-craver, CynthiaPublisher:McGraw-Hill Publishing Co.
Laboratory Manual For Human Anatomy & PhysiologyBiologyISBN:9781260159363Author:Martin, Terry R., Prentice-craver, CynthiaPublisher:McGraw-Hill Publishing Co. Inquiry Into Life (16th Edition)BiologyISBN:9781260231700Author:Sylvia S. Mader, Michael WindelspechtPublisher:McGraw Hill Education
Inquiry Into Life (16th Edition)BiologyISBN:9781260231700Author:Sylvia S. Mader, Michael WindelspechtPublisher:McGraw Hill Education





