
Concept explainers
(a)
Interpretation:
The
Concept introduction:
In mass spectroscopy, compounds can be identified on the basis of the mass of the compound. When the compound breaks into fragment then they can be distinguished from the other compounds. This technique is also used to differentiate the isotopes of compounds. In amino acids, three types of fragments are observed in low energy collisions are a, b and y ions. It is known as tandem mass spectrometry.
Answer to Problem 27.27P
The
Where N is asparagine, F is phenylalanine, E is glutamic acid, S is serine, G is glycine, K is lysine amino acid.
Explanation of Solution
In amino acids, b-type fragments appear due to an amino group or in other words charge is being carried by N-terminal. That is why it is also known as the N-terminus amino acid fragment. The b-type fragment is shown below.
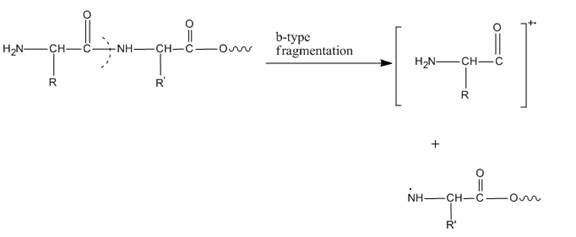
Figure 1
The given peptide is
The
(b)
Interpretation:
The
Concept introduction:
In mass spectroscopy, compounds can be identified on the basis of the mass of the compound. When the compound breaks into fragment then they can be distinguished from the other compounds. This technique is also used to differentiate the isotopes of compounds. In amino acids, three types of fragments are observed in low energy collisions are a, b and y ions. It is known as tandem mass spectrometry.
Answer to Problem 27.27P
The
Where N is asparagine, F is phenylalanine, E is glutamic acid, S is serine, G is glycine, K is lysine amino acid.
Explanation of Solution
In amino acids, y-type fragments appear due to a carboxyl group or in other words charge is being carried by C-terminal. That is why it is also known as the C-terminus amino acid fragment. The y-type fragment is shown below.
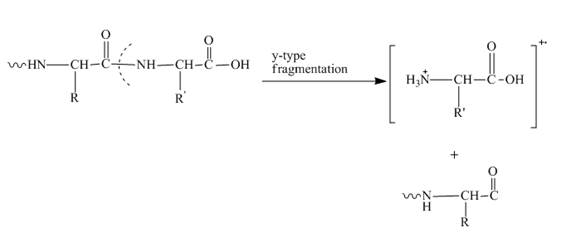
Figure 2
The given peptide is
The
Want to see more full solutions like this?
Chapter 27 Solutions
Organic Chemistry
- 22-61 Polyglutamic acid (a polypeptide chain made only of glutamic acid residues) has an a-helix conformation below pH 6.0 and a random-coil conformation above pH 6.0. What is the reason for this conformational change?arrow_forward22-49 Based on your knowledge of the chemical properties of amino acid side chains, suggest a substitution for leucine in the primary structure of a protein that would probably not change the character of the protein very much.arrow_forwardDescribe the tertiary and quaternarystructure of a protein.arrow_forward
- Q1)In the pH range 1.82 to 8.99, H2Arg+ is the principal form of arginine. Which is the second most prominent species at pH 6.0? At pH 5.0?Q2) (a) Draw the structure of the predominant form (principal species) of 1,3-dihydroxybenzene at pH 9.00 and at pH 11.00.(b) What is the second most prominent species at each pH?(c) Calculate the percentage in the major form at each pHarrow_forwardPara-aminobenzoic acid has two ionizable groups. One is a carboxylic acid that ionizes through loss of a proton thereby forming a charged para-aminobenzoic acid carboxylate anion, and the other is an amine connected to an aromatic (benzene) ring that ionizes by accepting an H+ to form a positively charged ammonium ion. The pk of the carboxylic acid is 4.9 and the pk of the amino group is 2.5. Assume that the intracellular pH within a bacterial cell is 7.0. N- H O-H H + H-N- H pK₂ = 2.5 Which of the following structures correctly depicts how most of the para-aminobenzoic acid is ionized at pH 7? A. B. O-H C. H H + H-N- H H D. H N- O-H H pK₂ = 4.9arrow_forwardPeptides can be separated using an ion-exchange column based on their isoelectric (pI) values. At which pH values would two different peptides, one with a pl of 5.6 and the other with a pl of 8.6, bind to a cation- and anion-exchange column? Each peptide may be capable of binding to each column at more than one pH value. anion-exchange column at pH = 4.0 pH = 6.5 pH = 10.1 cation-exchange column at = 4.0 pH pH = pH = 6.5 = 10.1 Answer Bank peptide B pl = 8.6 peptide A pl = 5.6arrow_forward
- The amino acid histidine has ionizable groups with pK₁ values of 1.8, 6.0, and 9.2, as shown. COOH H¸Ñ—CH CH₂ H 2 CH pH = COO™ H¸Ñ—CH 1.8 pk₁ CH₂ H lonizable -COOH = -COO- group CH COO™ H₂N-CH 6.0 pK₂2 CH₂ H -HisH -His N CH COO™ H₂N-CH 9.2 pk CH₂ H 2 —NH — —NH, CH A biochemist makes up 95 mL of a 0.13 M solution of histidine at a pH of 5.3. She then adds 60 mL of 0.10 M HCl. What is the pH of the resulting solution?arrow_forwardConsider the following peptide : Phe – Glu – Ser – Met and Val – Trp – Cys – Leu. Do these peptides have different net charges at (a) pH 1? (b) pH 7? Indicate the charges at both pH valuesarrow_forwardGiven below is the structure of a peptide with the name seryllysylaspartate. What is the amino acid at the C-terminus (or C-terminal amino acid) of this peptide? OH | CH, O || + H₂N-C | H O lysine Z-: serine + ŃH₂ T CH₂ T O aspartate CH₂ T CH₂ CH, O || N -C | O lysylaspartate H H C CH, O || C-O C-N C 1 H Harrow_forward
- . Describe the pH range of acceptable buffering behavior for the amino acids alanine, histidine, aspartic acid, and lysine.arrow_forwardComplete the following reaction: Naz-peptide -Peptide CA> ران CH CH3 CH3 NP₂ CH H' (hydrolysit) NA₂ V=0arrow_forwardAbout a decade ago, it was more common to use peptide mass mapping to identify a protein cut from a gel. Peptide mass mapping involves digesting the protein with trypsin and then measuring the peptide masses using MALDI-TOF mass spectrometry. The more superior method of identifying a protein from a gel is to digest the protein with trypsin and then use liquid chromatography to separate the peptides. The separated peptides are then electrosprayed into a tandem mass spectrometer. The MS-MS data is uploaded into a database such as MASCOT for protein identification. Explain why the LC-MS-MS method is superior to the peptide mass mapping method.arrow_forward
 Introduction to General, Organic and BiochemistryChemistryISBN:9781285869759Author:Frederick A. Bettelheim, William H. Brown, Mary K. Campbell, Shawn O. Farrell, Omar TorresPublisher:Cengage Learning
Introduction to General, Organic and BiochemistryChemistryISBN:9781285869759Author:Frederick A. Bettelheim, William H. Brown, Mary K. Campbell, Shawn O. Farrell, Omar TorresPublisher:Cengage Learning
